![]()
Sometimes I find it really frustrating that all we ever talk about is species (granted, I am very guilty of this too). Is it new? Where is it found? Where isn’t it found? Where do we put it on the tree of life? Does it need protection? Zzzzzzzzzzzzzz…
The question I wished we asked more is: what is this species doing? When I look at marine sediments under a microscope, my mind buzzes with the thought of millions of unknown genetic pathways all ecologically intertwined yet inherently isolated in each individual organism. In the old days (circa 2005 and boho-chic fashions), understanding gene expression was limited to single species—common model organisms—scrutinized in highly artificial settings (hmm, which gene do I want to knock out of this mouse today?). Oh, and you had to have previously sequenced the entire genome for said organism if you ever wanted a shot in hell at understanding your data.
Fast forward to the present, and researchers are embracing a new field: Metatrascriptomics. Try to say that three times fast. Transcriptomics of any variety is the study of RNA molecules (messenger RNAs, ribosomal RNAs, transcript RNAs and non-coding RNAs) present in a cell at any given time. By sequencing RNA molecules, we can get a snapshot of the genes being expressed in a cell, tissue, organism, or even whole community of organisms at a given place and time. These type of studies used to be carried out in a limited fashion using quantitative PCR (qPCR) or microarrays, but new sequencing technologies (454, Illumina) now allow us to sequence RNA at a much grander scale: we’re getting more sequences than ever (millions at a time) and can cover a huge taxonomic breadth (whole microbial communities).
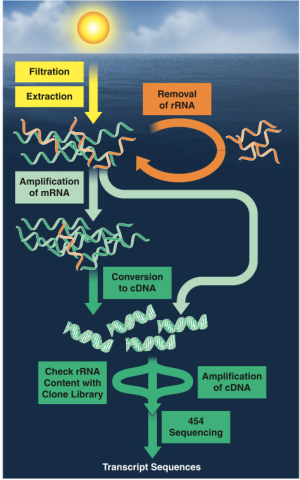
In the past few years, Metatranscriptomics has revealed some really neat stuff about marine environments, which Mary Ann Moran has summarized in a stellar review article. For example, Hewson et al. described the expression profile of a key cyanobacteria species (Crocosphaera watsonii) whose biology was poorly known but played a key role in nitrogen fixation in marine environments. In another study, Poretsky et al. analysed mRNA transcripts to describe the activity Pacific bacterioplankton communities across day and night. They found evidence that these organisms invest heavily in energy acquisition and metabolism when the sun is shining, but shift over to producing important molecular components (amino acids, vitamins, membranes) once immersed in darkness. How cool is that?

These new approaches are once again highlighting how little we know (when isn’t that the case in science?). It appears that any typical pelagic microbial transcriptome currently contains a huge proportion of previously unknown gene products—Gilbert et al. reported a whopping 91% of large gene families that appeared to be novel!
These studies just reinforce my (somewhat compulsive) desire to sequence everything in sight. That, and remind me to start carrying around an emergency pipetting kit, just in case.
References:
Gilbert, J., Field, D., Huang, Y., Edwards, R., Li, W., Gilna, P., & Joint, I. (2008). Detection of Large Numbers of Novel Sequences in the Metatranscriptomes of Complex Marine Microbial Communities PLoS ONE, 3 (8) DOI: 10.1371/journal.pone.0003042
Hewson, I., Poretsky, R., Beinart, R., White, A., Shi, T., Bench, S., Moisander, P., Paerl, R., Tripp, H., Montoya, J., Moran, M., & Zehr, J. (2009). In situ transcriptomic analysis of the globally important keystone N2-fixing taxon Crocosphaera watsonii The ISME Journal, 3 (5), 618-631 DOI: 10.1038/ismej.2009.8
Moran, M.A. (2009). Metatranscriptomics: Eavesdropping on complex microbial communities Microbe, 4 (7), 329-335 Open-Access PDF
Poretsky, R., Hewson, I., Sun, S., Allen, A., Zehr, J., & Moran, M. (2009). Comparative day/night metatranscriptomic analysis of microbial communities in the North Pacific subtropical gyre Environmental Microbiology, 11 (6), 1358-1375 DOI: 10.1111/j.1462-2920.2008.01863.x





One days we’ll have cheap, portable sequencers that we can pass out to citizen scientists. That will be an awesome day.
What a great methodology. I can imagine this would provide some really interesting insights into genetics and ecology. Good post thanks!