![]() Sometimes I am stunned by the vastness of the internet, as well as the brief 15-nanoseconds of fame that go along with most of its content. The other day I discovered the ‘Charlie the Unicorn’ videos on YouTube, after (ironically?) having a conversation with a real three-dimensional human.
Sometimes I am stunned by the vastness of the internet, as well as the brief 15-nanoseconds of fame that go along with most of its content. The other day I discovered the ‘Charlie the Unicorn’ videos on YouTube, after (ironically?) having a conversation with a real three-dimensional human.
I was excited by this hilarity and went to whip up a post, only to discover that this video had already been posed by Dr M. In April, 2009. After being publically berated by Kevin, I remembered where I was back in 2009—locked in an apartment in London, in a self-inflicted thesis-writing exile, too stressed to even go grocery shopping. I developed temporary synesthesia from staring at DNA alignments for 3 months on end; the A’s, T’s, C’s and G’s in black text would pop into color as soon as my brain recognized them. Tweaking alignments became an obsessive hobby, akin to Frogger or PacMan—so simple, yet never boring and always unfinished.
Although much of 2009 passed in a hazy blur, I remember remember the 5th of November when I finally submitted my Ph.D. thesis. (yes, I timed my submission date to coincide with Guy Fawkes’ Day because I am that kick-ass).
The last two papers from my thesis were finally published a few months ago in BMC Evolutionary Biology, but they’re a dense read (unless the phrase “likelihood search of tree space with bootstrapping under a gamma distribution” gets you all hot and bothered)
My Ph.D. research was pretty cool because:
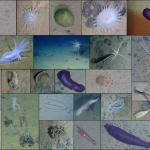
- It was the first publication to place deep-sea nematodes on the Tree of Life using DNA sequences
- I focused on a specific (but typically neglected) group of predatory worms called the Enoplida. By the time I finished sequencing, my data resulted in a 15-fold increase in the number of Enoplid sequences in GenBank.
The end result was a pretty picture like this (where red are deep-sea taxa and blue are shallow water taxa):

What did my work show?
Deep-sea nematodes have a complex evolutionary history, with gene sequences suggesting much shuffling between shallow and deep habitats. Nematodes didn’t invade the deep sea at one point in time—instead, movement across depth gradients happened in many different taxa at various points in time.
There is a high diversity within deep-sea nematode communities—for example, I found what looked like three distinct species within one GENUS in a sub-Antarctic site. But you wouldn’t know this by looks alone—some families had hugely divergent gene sequences despite very similar anatomical features. Molecular evidence also suggested the existence of closely related (putatively cosmopolitan) species complexes in the deep-sea; Enoplid nematodes didn’t show any patterns related to depth or geographic location.

Finally, my work tried to figure out where the branch of Enoplid nematodes had grown out on the Tree of Nematodes. In short, we couldn’t get a clear answer. Building the Tree of Life is like choosing your spectacles—sometimes you want to look really close (so you need reading glasses, e.g. rapidly evolving genes) but other times you’re straining to see what’s in the distance (and you need strong lenses, e.g. conserved, slowly evolving genes). But even though you might have the Hubble Telescope to peer at far-distant objects, some galaxies still lie beyond the telescope’s resolution and appear fuzzy. Same thing with the conserved genes that encode the small ribosomal subunit (18S rRNA, in our case)—we confirmed that Enoplids are a divergent, old lineage of nematodes, but exactly where they branched off is still a bit fuzzy.
References:
Bik, H., Thomas, W., Lunt, D., & Lambshead, P. (2010). Low endemism, continued deep-shallow interchanges, and evidence for cosmopolitan distributions in free-living marine nematodes (order Enoplida) BMC Evolutionary Biology, 10 (1) DOI: 10.1186/1471-2148-10-389
Bik, H., Lambshead, P., Thomas, W., & Lunt, D. (2010). Moving towards a complete molecular framework of the Nematoda: a focus on the Enoplida and early-branching clades BMC Evolutionary Biology, 10 (1) DOI: 10.1186/1471-2148-10-353




Cool work Holly! Did you know if the functional ecology of deep-sea lineages reflects their shallower ancestors? Like, did they inhabit similar niches in the deep, follow shallow preferred prey deeper or is it a mixed bag of generalist predators shallow and deep?