Ironically enough, I was at a meeting about oil spills when the Macondo well blew. The “Natural Resource Damage Assessment (NRDA) in Arctic waters” workshop brought scientists and industry contacts together to discuss the challenges and consequences of petroleum-related accidents in fragile polar habitats. I remember the BP executives had to step out to deal with a small “issue” one evening. By morning, they had disappeared entirely.
When all the shiz went down in the Gulf of Mexico, yours truly and collaborators had their nose to the grind, madly running around collecting samples and spending late nights in the lab listening to the hum of PCR machines. We were awarded an NSF RAPID grant in August 2010 to use parallel taxonomic and high-throughput sequencing approaches to characterize the impacts of the Deepwater Horizon oil spill on microscopic eukaryote communities inhabiting marine sediments. In English: we used both DNA and old skool microscopy to compare species living on beaches before and after the oil spill. This grant funded some hardcore sampling trips (September 2010, where I went on a boat and drove 1700+ miles along the Gulf Coast in one week, and March 2011 when we returned to sample sites a year after the oil spill) and a kickass undergraduate workshop about the “Bioinformatics of Biodiversity” which may or may not have involved YouTube Karaoke sessions and a randomly acquired cowboy hat.
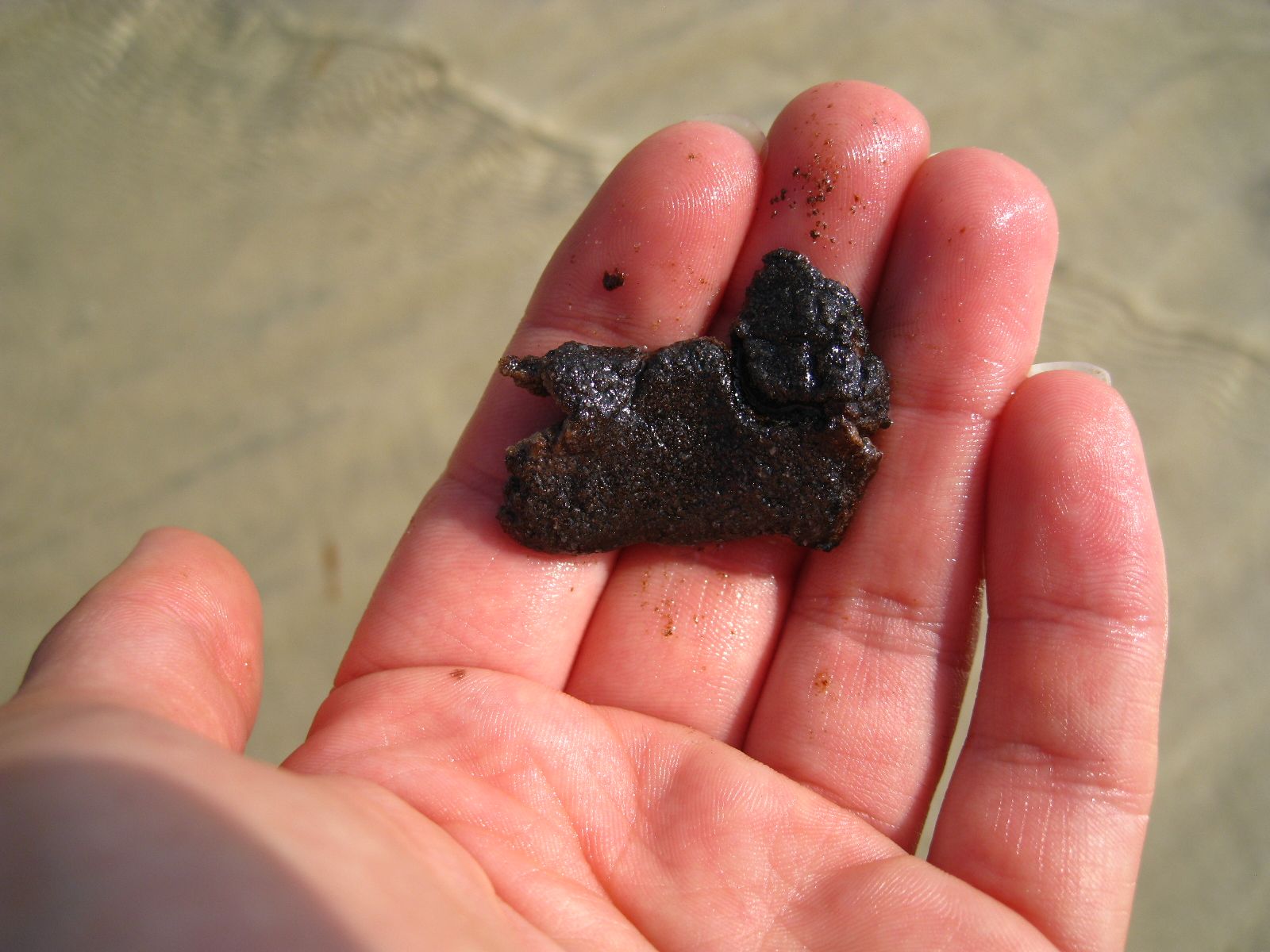

So today, I’m pleased to announce that the first (of hopefully many) papers from this project has officially been published in PLoS One. For our first manuscript, we focused on a set of pre- and post-spill samples collected around Dapuhin Island, Alabama in May and September 2010, respectively. Our pre-spill samples represented our baseline, collected by our collaborator Ken Halanych at Auburn University days after the oil started gushing (his team quickly drove down to the coast before any of it had come close to the shore). Post-spill samples were collected 4 months later, after sheets of sticky crude were pitched ashore during a summertime of heavy beach oiling and BP’s cleanup efforts doggedly wiped away the blackness. We also included another post-spill sample site from Grand Isle, Louisiana, where I suspected that the the state of the beach would produce some intriguing data.
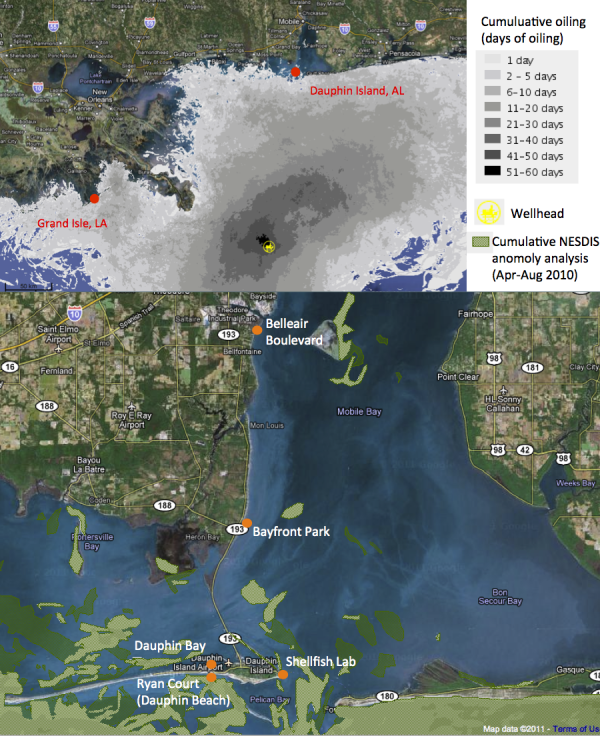
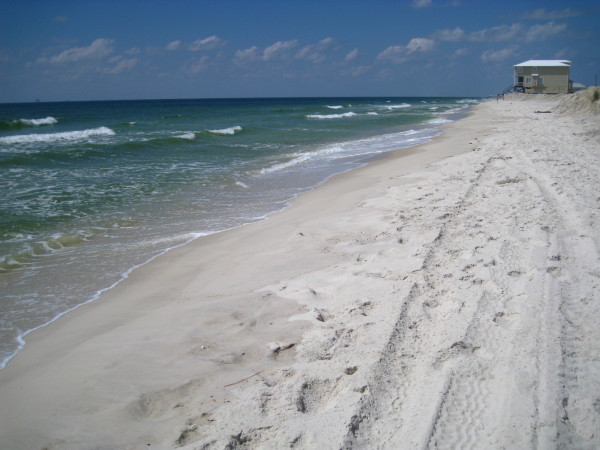
Before I describe our results, I’ll show you what the beaches looked like at the time of post-spill sampling. In September 2010, Dauphin Island was pretty serene: quiet, yes, because tourists were eschewing the region, but the shoreline itself showed little evidence of the BP fiasco. If you looked closely (which I certainly did), you could find little splotches of oil, an isolated tarball, or a buried “dirty” layer of sand. But if you had spent the past year on a media blackout, never heard of Deepwater Horizon, you would think that the Alabama coast looked pretty ordinary.

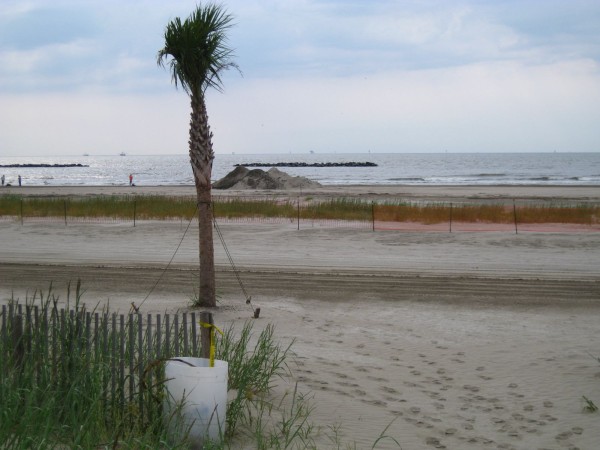
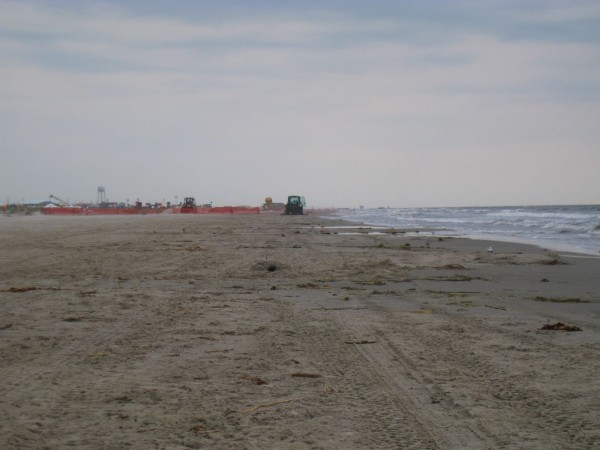
Grand Isle was a scene from another planet. A far cry from the parallel, reestablished tranquility on other Gulf shores. Grand Isle was a beach that was clearly impacted at the time of sampling. In fact, I specifically made a 6-hour detour to this site after hearing local news reports lamenting those tarnished Louisiana shores. Upon my arrival, I found the beach to be so impacted that I was hardly allowed near the sea. The ranger on duty turned me away at the local park. Fluorescent orange mesh blocked my attempts to cross the dunes. When I finally found a gap that led me to water, I made haste for fear of being chased away. There was heavy machinery humming up and down the shore (the park ranger noted that beach access was restricted for safety concerns), and piles of oiled sand awaiting a mechanical cleanse.


At each site we collected replicate sediment cores for DNA work (frozen immediately on dry ice) and taxonomic analysis (archived in 4% formalin, which is better for preserving morphology). The sequencing itself was a piece of cake (although the bioinformatics, not so much). I carried out standard environmental DNA protocols at the Hubbard Center for Genome Studies at the University of New Hampshire, where I was previously working as a postdoc under Kelley Thomas (senior author on our paper). First we separated the microbial eukaryote species from the sediment by suspending them in water and concentrating these organisms on a 45um sieve. Next we broke open all the cells using a beadbeather: think “Will it Blend” with ball bearings and soft tissue. Nothing stays intact. Finally, we used the Polymerase Chain Reaction (PCR) to broadly amplify two different regions of the 18S rRNA gene from the entire biological community present at each sample site. PCR amplicons were sent off for 454 sequencing, and we waited. In the meantime, Jo Sharma at UTSA was spending long hours at the microscope carrying out taxonomic identifications for nematodes at each site being sequenced.
I’ll pause for a moment here to offer more context. When I say we are studying “microbial eukaryote” species, I’m talking about puny things with a body size <1mm. You know, the ones no one cares about. And the ones I happen to be obsessed with (they’re so much more interesting than dolphins). We’re talking about taxonomic groups like meiofaunal metazoans (e.g. Nematoda, Platyhelminthes, Gastrotricha and Kinorhyncha, etc.), microbial representatives of fungi and deep protist lineages (Alveolata, Rhizaria, Amoebozoa, algal taxa in the Chlorophyta and Rhodophyta, etc.), and eggs and juvenile stages of some larger metazoan species. The reason why we chose to focus on these groups is precisely because they tend to be ignored. Most of the awesome genomic investigations only look at Bacteria and Archaea. But small eukaryotes are equally ubiquitous as their non-nucleated counterparts, and in marine ecosystems they play key roles as decomposers, predators, producers and parasites–yet we know little about their biology, ecology and diversity. By describing species changes in the Gulf of Mexico, we wanted to infer something about the potential for large-scale or long-term repercussions for Gulf ecosystems.
Sorry to keep you waiting–lets get down to the juicy stuff. Our results were pretty dramatic. After analyzing 1.2 million DNA sequences alongside nematode taxonomy, we found shockingly significant shifts in microbial communities between pre- and post-spill sites.
The first thing we saw was a stark shift in the diversity and abundance of taxa between pre- and post-spill sites. Pre-spill sites showed a typical marine community: dominated by nematodes, but containing a mishmash of other taxa such as arthropods, polychaetes, protists, algae, and fungi. Post-spill sites, in contrast, were almost exclusively dominated by a few species of fungi, with a spattering of some other metazoan species.

After looking at the charts summarizing the overall taxonomic assemblages, we moved on to ecological analyses such as Unifrac. We built a phylogenetic tree with our DNA sequences, computed some metrics about the branching topology, and got an overarching indication of how similar our samples sites were at the community level (e.g. for all species sequenced at each site).
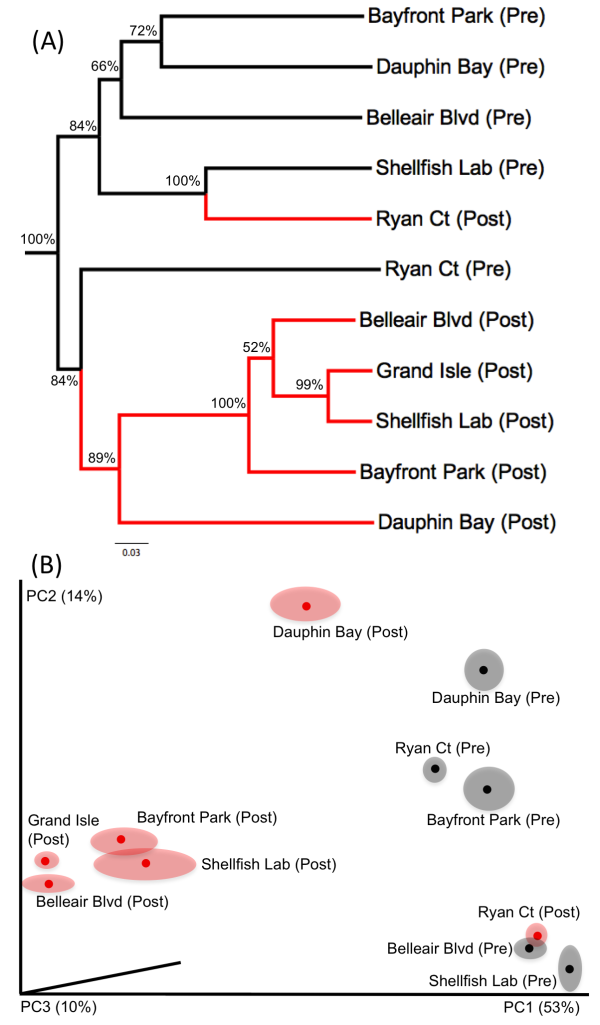
We also did something similar with taxonomic data, using the Bray-Curtis similarity metric to analyze the list of visually-identified nematode species which were present (or not) across our sample sites.

Both our DNA and taxonomic analyses were relaying the same story: our samples clustered together according to pre- and post-spill time points: the before/after communities at the SAME site weren’t closely related to each other. Principal Coordinates Analysis (PCoA, Unifrac figure B, above) also underlined biodiversity distinctions across pre-spill sites. Even though pre-spill sites were characterized by nematode dominance, it wasn’t the same group of nematode species present at every site. In contrast, post-spill sites converged towards a similar community structure–these trends were likely driven by oil-associated fungal taxa that were common across post-spill sites.
You’ll also note that we observed a few outlier sites; Ryan Court (a sandy beach in front of residential property, on the Gulf coast of Dauphin Island) and Dauphin Bay (an inlet on the opposite site of the island, facing the Alabama mainland). Although we did observe community shifts in these post-spill samples, the shifts weren’t characterized by the typical fungal dominance seen at other sites. We think this has something to do with the geography and human-mediated cleanup efforts. To protect residents, Ryan Court had waterborne barriers going up and down during the heaviest oiling–this might have mitigated the worst effects in the sediment perhaps, preventing a shift to fungal dominance. Oil also might not have penetrated the inland Dauphin Bay site very well, since it was inherently sheltered by its location and some nearby marshland on Dauphin Island.
For me, the most convincing evidence of oil impacts was the data from Grand Isle. That 5-hour drive (and accompanying True Blood soundtrack) was the best sampling decision I made. Although I was pretty scared of Vampires when I was driving back through Louisiana that night. The beach was unarguably facing heavy oil impacts when I took samples. DNA analysis showed that the fungal-dominated post-spill assemblage in LA contained the same taxa as the Shellfish Lab, Dauphin Island community. Same putative species (Operational Taxonomic Units, a.k.a. OTUs), found 250 miles apart. To the casual observer, the scene at Dauphin Island didn’t look anything like Grand Isle. But thanks to the deep insight afforded by high-throughput sequencing, we were able to capture a snapshot of post-spill microbial assemblages that was highly indicative of environmental disturbance.
So we started looking closer at the data. Looking within phylogenetic tree topologies, I manually examined what taxa were most closely related to our fungal OTUs. Evolutionary relationships seemed to hint that post-spill fungi could survive using environmental hydrocarbons as an energy source:
Two distinct fungal community structures were recovered at post-spill sites: one assemblage dominated by Cladosporium OTUs (recovered at Shellfish Lab and Grand Isle), showing a close relationship to C. cladosporioides sequences in phylogenetic topologies, and a second assemblage dominated by OTUs in the fungal genus Alternaria (Belleair Blvd and Bayfront Park). Fungal taxon dominance may be dictated by the physical marine environment; Alternaria OTUs dominated in brackish Mobile Bay, while Cladosporium was recovered in higher-salinity sediments on the outer shores of Dauphin Island. These highly dominant post-spill OTUs appear as rare taxa in diverse pre-spill fungal assemblages, suggesting that oil-induced environmental stress may have favoured the rise of resilient, opportunistic species (able to capitalize on the large input of new resources). Although the diversity and ecological role of marine fungi is not well understood, previous evidence suggests that observed fungal assemblages denote a signature of crude oil in Gulf sediments. Cladosporium contains ubiquitous, opportunistic species that can extensively utilize hydrocarbon compounds and thrive in hostile, polluted conditions that appear to be intolerable for other marine fungi [9,10]. Compared to many other fungi, marine Altenaria demonstrate increased activity of lignocellulose-degrading enzymes [11] that have been implicated in breakdown of industrial toxins [12,13]. In addition to these dominant OTUs, we recovered a variety of fungi at post-spill sites (including OTUs phylogenetically related to Apergillus, Acremonium, Acarospora, Rhodocollybia, and Rhizopus) that rarely comprised a significant component of pre-spill fungal communities. A number of these marine groups have also been shown to metabolize hydrocarbon compounds [14,15]. (Bik et al. 2012)
This paper has been a long time coming, and I’ve been dying to blog about it for the better part of a year. We went through the rounds and rejections at several top-tier journals before a lengthy review process at PLoS ONE, so I’m now pleased that these results are finally seeing the light of day. Our work in the Gulf of Mexico is still ongoing — unfortunately this was one study that raised a hell of a lot more questions than answers. We’ve continued to collect post-spill samples at regular intervals (including the samples I collected one-year after the original pre-spill samples). We want to figure out if the community shifts we saw in this study were really due to oil (as suggested by the dominance of oil-associated fungal taxa), mechanical beach cleanup efforts (which may have physically damaged and killed fragile microbial species), or whether they might be influenced by seasonal and temporal variation in the Gulf region (a topic where there isn’t much existing data). Another motivation is to study the longevity of these patterns–additional sampling time points will allow us to track the post-spill recovery, or lack thereof, of microbial eukaryote communities. Will assemblages begin to resemble pre-spill communities again, or will these beaches remain depauperate and/or be replenished with a different set of fauna? The post-spill fungi are cool and intriguing too. Were they thriving in oiled beach sands, or just weakly persisting after other species were killed off? Transcriptomics (studying gene expression by sequencing mRNA) will help us to answer this question and determine the species that were alive and kicking at the time of sampling. We’re also going to use random, shotgun sequencing to look deeply into the genomes of sparsely-populated, fungal-dominated beaches at sites such as Grand Isle: if post-spill species are eating hydrocarbon compounds, perhaps their genetic machinery will give an indication of the metabolic pathways that enable them to use oil as an energy source.
So really, this first paper is just a prelude. The main act will be as grand as Beethoven’s 5th.
Reference: Bik, H.M., Halanych, K.M., Sharma, J. & Thomas, W.K. (2012) Dramatic shifts in benthic microbial eukaryote communities following the Deepwater Horizon oil spill, PLoS ONE http://dx.plos.org/10.1371/journal.pone.0038550
Share the post "Dramatic impacts on beach microbial communities following the Deepwater Horizon oil spill"



The language of this blog was way over my head for the most part. I get the sense that there HAS been a radical change in the types and quantity of these small animals. (think bottom of the food chain) I was there in June 2010 as part of the Response effort and KNEW that from the volumeof oil and dispersant being dumped into the Gulf that this would be DECADES before we know the extent of the damage. I believe that we are only now seeing the beginning of the death of the Gulf as a viable ecosystem. ANd I hold BP AND our gov’t directly responsible for this debacle.
WOW Dr Bik. Amazing science and equally killer blog post. You continue to inspire.
Q: Are you repeating this sampling? I wonder to what extent this is a short-term transitory community versus a long-term state-shift.
A friend linked this to me. I’m on the responder side.
I’ll just have to add that yes, this is indeed an expected behavior as the point of many response techniques is to enhance biodegredation of hydrocarbons. Thus, it is expected that the microfauna can will shift towards a community that is high in components which process hydrocarbons into bioavailable materials. A radical change, in fact.
But, this bioavailability also should protect other microfauna/flora from acute effects of the pollution, even with local mortality, as rapid degradation assists in detoxification which then results in quicker recolonization.
It’s a shame that pre-spill data isn’t available. It would greatly assist in the NRDA process and determining lowered upper trophic level biovailability and corresponding lowered economic outputs in harvestable stock.
Also interesting will be the following seasonal effects as recolonization of surfaces happen, in the next 20 years or so. From the available historical data, microbiota effects are transitory over that timeframe. It would be interesting to reconfirm it in this case as well.
Robert, dispersants are there to increase bioavailability. That’s their purpose. It can cause localized short term chronic effects. Inherent in that is some short term increase in polyaromatic hydrocarbon metabolisis. The key is, monitoring to ensure that sufficient stock remains to recolonize, and that entire stocks aren’t reduced permanently because of acute mortality.
Also clouding this data is the ongoing eutrification of the gulf from upstream sources, and an ongoing permanent shift in food web configuration due to climactic and human harvesting factors …
This is an excellent paper and extremely valuable to us in trying to get implemented effective, non-toxic clean-up solutions for contaminated waters. As there is significant documentation showing that nowhere near 100% cessation of the flow of oil was ever achieved, as well as the continuing, covert use of the toxic chemical dispersant Corexit to sink and/or keep fresh oil beneath the surface waters, we sincerely look forward to your on-going research as this disaster unfolds and its impact continues to impair and seriously alter the Gulf’s delicate eco systems. http://www.TheEarthOrganization.org.
The fungi have taken over due to the depletion of O2 in all zones of the ocean eco system. The dispersants contained 2 butoxy ethanol and DOSS which prevents the breakdown of hydrocarbons, and causes the dispersant and toxicity to linger in the water column, sea bed and beaches. Numerous reports show that just a couple of inches under the sand there is oil and tar balls. The fact that these scientist state the micro flora has changed is obvious from the chemicals used in the spill, and their is evidence the spill is ongoing, which would be cause for some speculation of this report. There is still large amounts of tar balls and sheens seen on the ocean so with this continued onslaught of toxic dispersants and oil, the eco system is giving rise to species that are not particularly wanted, due to their inherent problems. This is more evidence that dispersants should not be used on spills.